Chapter 12 Regression analysis with many variables
12.1 Introduction
The standard regression model consists of fitting an equation of the form
\[y = \beta_{0} + \beta_{1}x_{1} + \beta_{2}x_{2} + \ldots + \beta_{p}x_{p}\] to a data set including a response/dependent variable \(y\) and a number of explanatory/independent variables \(x_1,\ldots,x_p\). However, when faced with many explanatory variables or more variables than samples in the data set, straightforward multiple regression struggles or simply fails. In particular the standard model is unsatisfactory when:
- \(p\) is large relative to \(n\),
- the explanatory variables \(x_i\) are highly correlated,
- and directly fails when \(p\geq n\).
We here briefly describe and illustrate two popular approaches to get around these difficulties:
- Principal Component Regression (PCR).
- Partial Least Squares (PLS).
Not surprisingly at this point of the course, the idea consists of projecting the data onto low dimensions.
12.2 PC regression of the NIR data set
The data set consists of near-infrared reflectance (NIR) spectra from 39 samples of wheat flour which account for the intensity of reflected light measured (optical density, OD) at 68 wavelengths (in increasing order). Additionally, sample protein content was assessed by wet chemistry.
Protein content and moisture are in the first columns and the NIR spectra along wavelengths are in the remaining columns. The objective is to fit a regression model to predict protein content from NIR spectra. The problem is that ordinary regression (say using the lm function in R) will fail because the data matrix has more columns than rows. Even if we had 80 or 90 samples there are still some additional estimation problems. Firstly, how do we select the important wavelengths from among so many? Secondly, if we select many wavelengths there is a strong chance of over-fitting. Finally, the ODs are highly correlated with each other (you can check this out using cor on a subset of them), which makes variable selection even more difficult.
Before proceeding we obtain a plot of the NIR spectra. The data values are columns 3 to 70 of the data.frame.
wv <- 1:68 # Range of wavelengths
# Extract spectra ODs and transpose to arrange them by rows (required by matplot)
OD <- t(nir[, 3:70])
matplot(wv, OD, type = "l", xlab = "Wavelength", ylab = "Optical Density (OD)",
main = "39 NIR Spectra")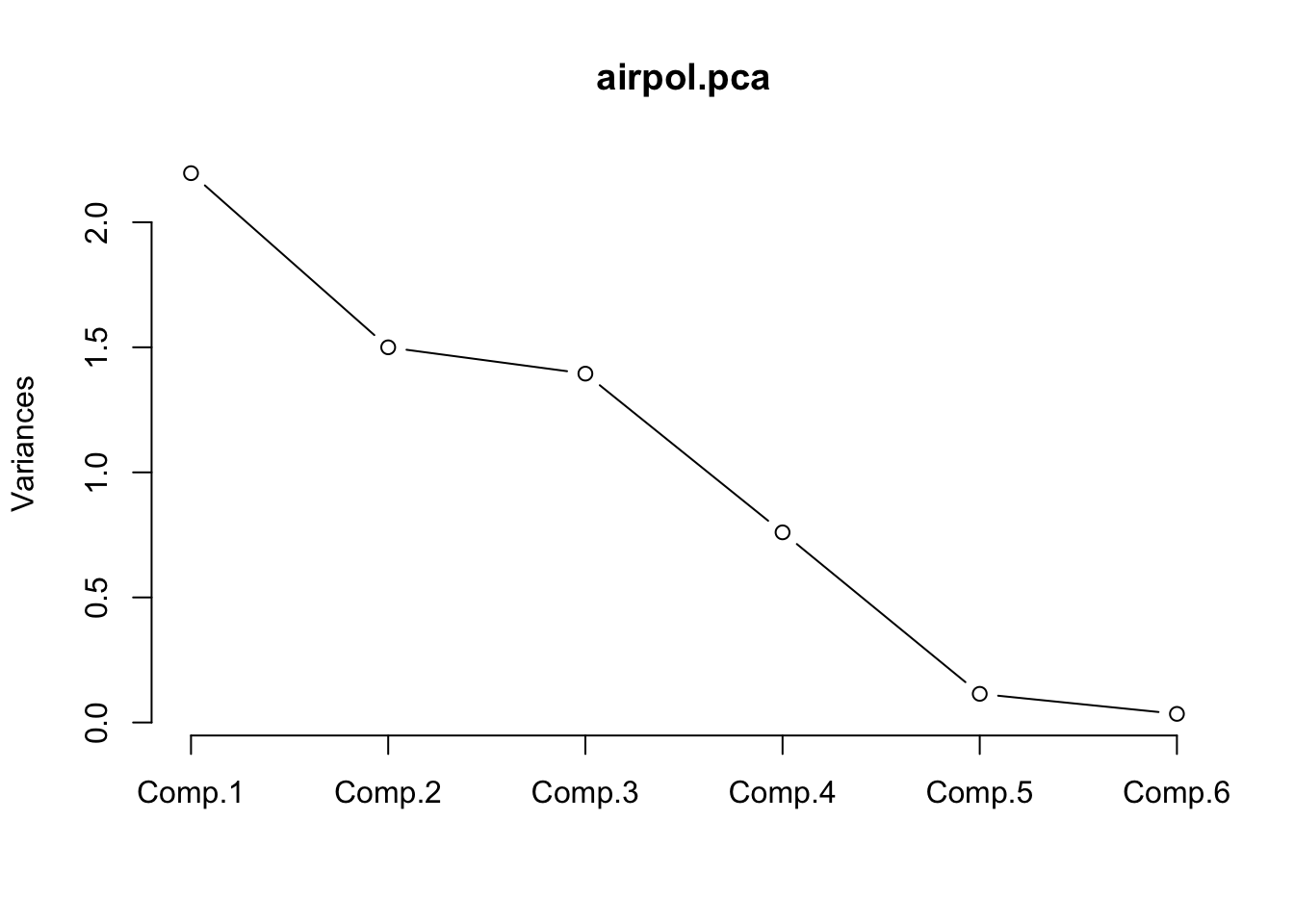
We can apply PCA on the explanatory variables \(\mathbf X\) and fit a regression equation of the form
\[\mbox{Protein} = \beta_{0} + \beta_{(1)}PC_{(1)} + \beta_{(2)}PC_{(2)} + \ldots + \beta_{(k)}PC_{(k)}\]
where \(PC_{(i)}\) refers to the \(i^{th}\) selected principal component from \(\mathbf X\).
But which PCs we choose? It is a good idea to select those which have the strongest correlation with the response variable, protein in this case. Note that these are not necessarily the first PCs (those explaining most variability in \(\mathbf X\)).
We derive the PCs and select those which have the strongest correlation with protein.
# Principal component analysis
nir.pca <- prcomp(nir[, 3:70])
# Consider e.g. the first 8 PCs
scores <- predict(nir.pca)[, 1:8]
# Extract loadings from PCA object
loadings <- nir.pca$rotation The following computes the correlation between the response variables and the PCs.
PC1 PC2 PC3 PC4 PC5 PC6 PC7 PC8
protein 0.6988 0.0925 -0.1891 0.6677 -0.1079 -0.0095 0.0280 0.0018
moisture 0.1634 -0.9735 -0.0908 -0.0444 -0.0625 -0.0216 -0.0092 0.0385Notice how it is only the first, third perhaps, and fourth PC which are correlated with protein. This is interesting given the % variance accounted for by the PCs (check for this using summary on nir.pca).
We now fit ordinary regression of protein on the selected PCs (\(1^{st}\), \(3^{rd}\) and \(4^{th}\) PCs).
Call:
lm(formula = nir[, 1] ~ scores[, c(1, 3, 4)])
Residuals:
Min 1Q Median 3Q Max
-0.64275 -0.22178 0.09703 0.23163 0.56511
Coefficients:
Estimate Std. Error t value Pr(>|t|)
(Intercept) 10.80000 0.05243 205.998 <2e-16 ***
scores[, c(1, 3, 4)]PC1 6.89984 0.28960 23.825 <2e-16 ***
scores[, c(1, 3, 4)]PC3 -39.69301 6.15733 -6.446 2e-07 ***
scores[, c(1, 3, 4)]PC4 192.41470 8.45166 22.767 <2e-16 ***
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
Residual standard error: 0.3274 on 35 degrees of freedom
Multiple R-squared: 0.9699, Adjusted R-squared: 0.9673
F-statistic: 375.8 on 3 and 35 DF, p-value: < 2.2e-16How good is the fit? We can look at the \(R^2\) of the model and plot fitted versus observed protein values.
plot(nir[, 1], pcr$fitted.values, xlab = "Observed", ylab = "Fitted")
abline(0, 1) # Add reference line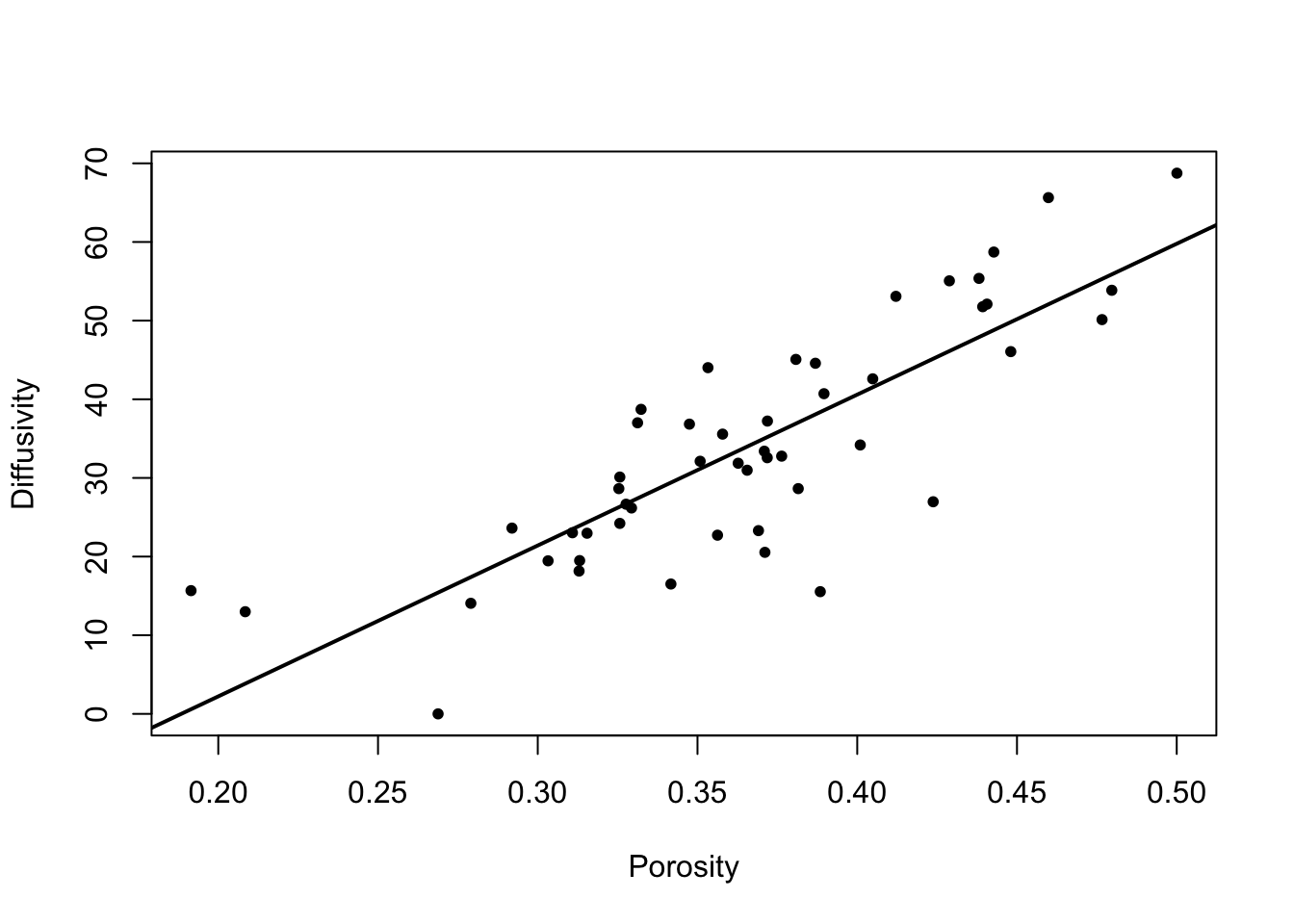
The coefficient of determination \(R^2=0.9699\) is very high and the plot of observed versus fitted protein values shows high agreement. Hence, measuring NIR spectra and processing them through PCR seems to be a reliably approach to predict protein content in wheat flour as an alternative to direct measurement.
It is also possible to express this regression in terms of the original wavelengths. We have
\[\mbox{Protein} = \beta_{0} + \beta_{(1)}PC_{(1)} + \ldots + \beta_{(4)}PC_{(4)}\]
where each PC can be expressed as \(PC_{(i)}=l_{1(i)}x_{1}+\ldots+l_{68(i)}x_{68}\). We can simply substitute and gather together the terms for each wavelength \(x\) to express the PCR models as
\[\mbox{Protein} = \beta_{0} + \delta_{1}x_{1} + \ldots + \delta_{68}x_{68}\]
A plot of these coefficients illustrates the relative weight of each wavelength for the prediction of protein.
beta <- pcr$coefficients
pcrmodel <- beta[2] * loadings[, 1] + beta[3] * loadings[, 3] + beta[4] * loadings[, 4]
plot(wv, pcrmodel, type = "l", ylab = "PCR Coeffs", main = "Protein, PCs 1,3,4")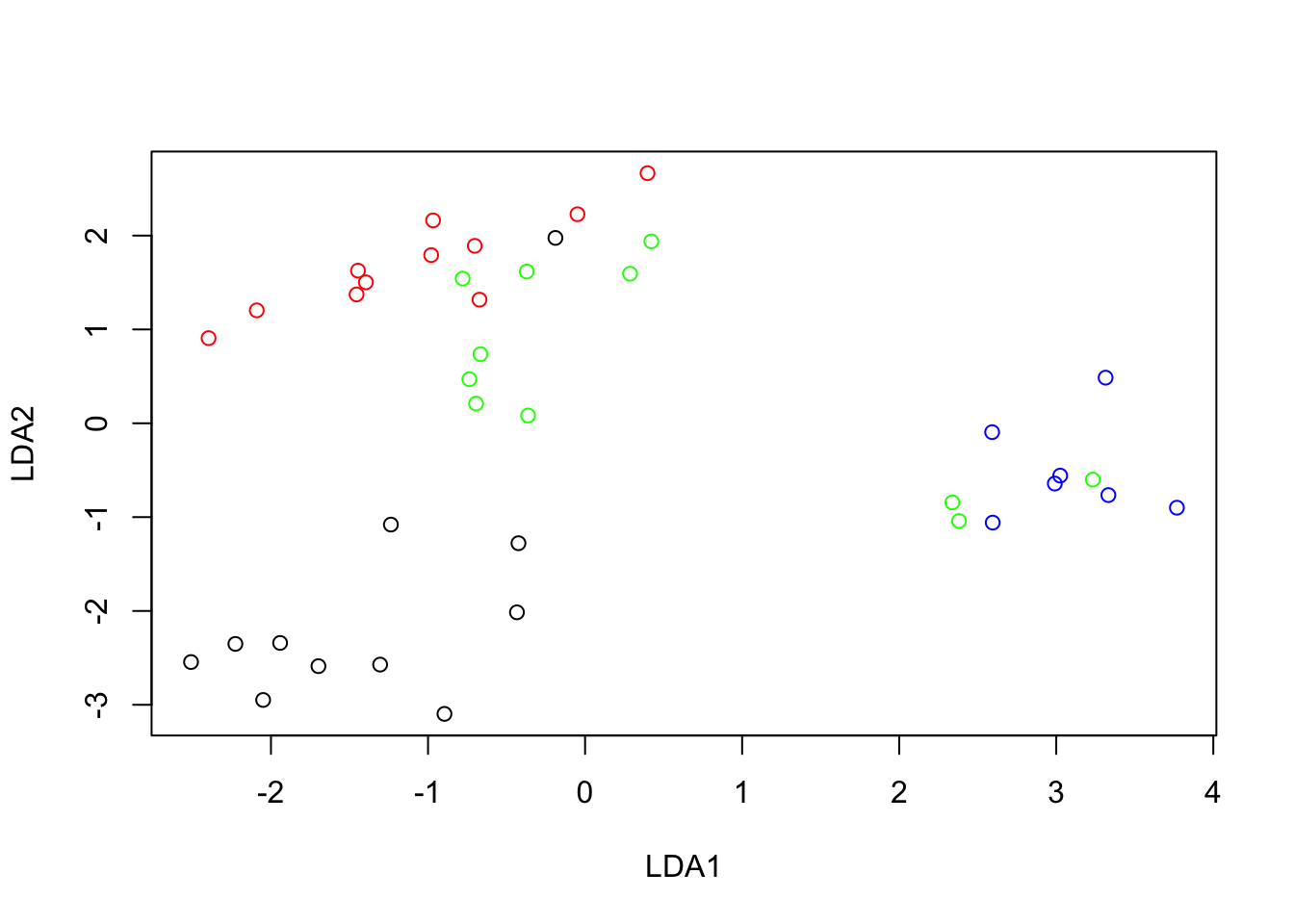
12.3 PLS regression of the NIR data set
Partial least squares is an alternative to principal component regression and is very popular in the field of chemometrics. PLS addresses the same problem as PCR. Given a large number of independent variables \(x_i\), often highly correlated, the purpose is to find a small number of linear combinations of the independent variables which are best predictors of the dependent variable \(y\).
There are few computational formulations of the PLS algorithm but, in brief, PLS aims to find combinations
\[\mathbf{\lambda'x} = \lambda_1x_1+ \lambda_2x_2 + \ldots + \lambda_px_p\]
which maximise
\[\mbox{Corr}^{2}(\mbox{$y$},\mathbf{\lambda'x})\mbox{Var}(\mathbf{\lambda'x}).\]
Some distinctive features are:
- PLS uses both the values of \(y\) and the \(x_i\) to find the latent components (often called factors).
- PCR uses \(\mathbf{X}\) only to find the new components (PCs), then makes the link to \(y\) by ordinary regression.
- PLS factors and scores are interpreted as for PCs (PLS factors also uncorrelated).
- For regression, PLS selects the first \(k\) PLS factors based on model performance (\(R^2\) and root mean squared error, RMSE, commonly).
- PLS generally uses 1 or 2 fewer terms than PCR to reach comparable goodness of fit.
We use the pls package to fit a PLS model to the NIR data set analogous to the previous one using PCR.
library(pls)
wv <- data.frame(nir[, 3:70])
pls <- plsr(nir$protein ~ ., ncomp = 6,data = wv, validation = "CV")
summary(pls)Data: X dimension: 39 68
Y dimension: 39 1
Fit method: kernelpls
Number of components considered: 6
VALIDATION: RMSEP
Cross-validated using 10 random segments.
(Intercept) 1 comps 2 comps 3 comps 4 comps 5 comps 6 comps
CV 1.835 1.347 0.9859 0.3670 0.2409 0.2364 0.2456
adjCV 1.835 1.343 0.9982 0.3623 0.2383 0.2331 0.2414
TRAINING: % variance explained
1 comps 2 comps 3 comps 4 comps 5 comps 6 comps
X 98.60 99.18 99.72 99.92 99.97 99.98
nir$protein 48.97 78.17 97.28 98.85 99.06 99.24The ncomp argument controls up to how many latent PLS factors or components we want to consider. As discussed in previous chapters, cross-validation is convenient here as well to have a more realistic estimate of the goodness of the fit. The cross-validated Root Mean Squared Error of Prediction (RMSEP) suggests that 4 PLS components would be enough. The \(R^2\) for the successive number of PLS components is obtained by
1 comps 2 comps 3 comps 4 comps 5 comps 6 comps
train 0.4897 0.7817 0.9728 0.9885 0.9906 0.9924
CV 0.4324 0.6958 0.9578 0.9818 0.9825 0.9811Goodness of fit using 6 PLS components using a observed versus predicted plot.
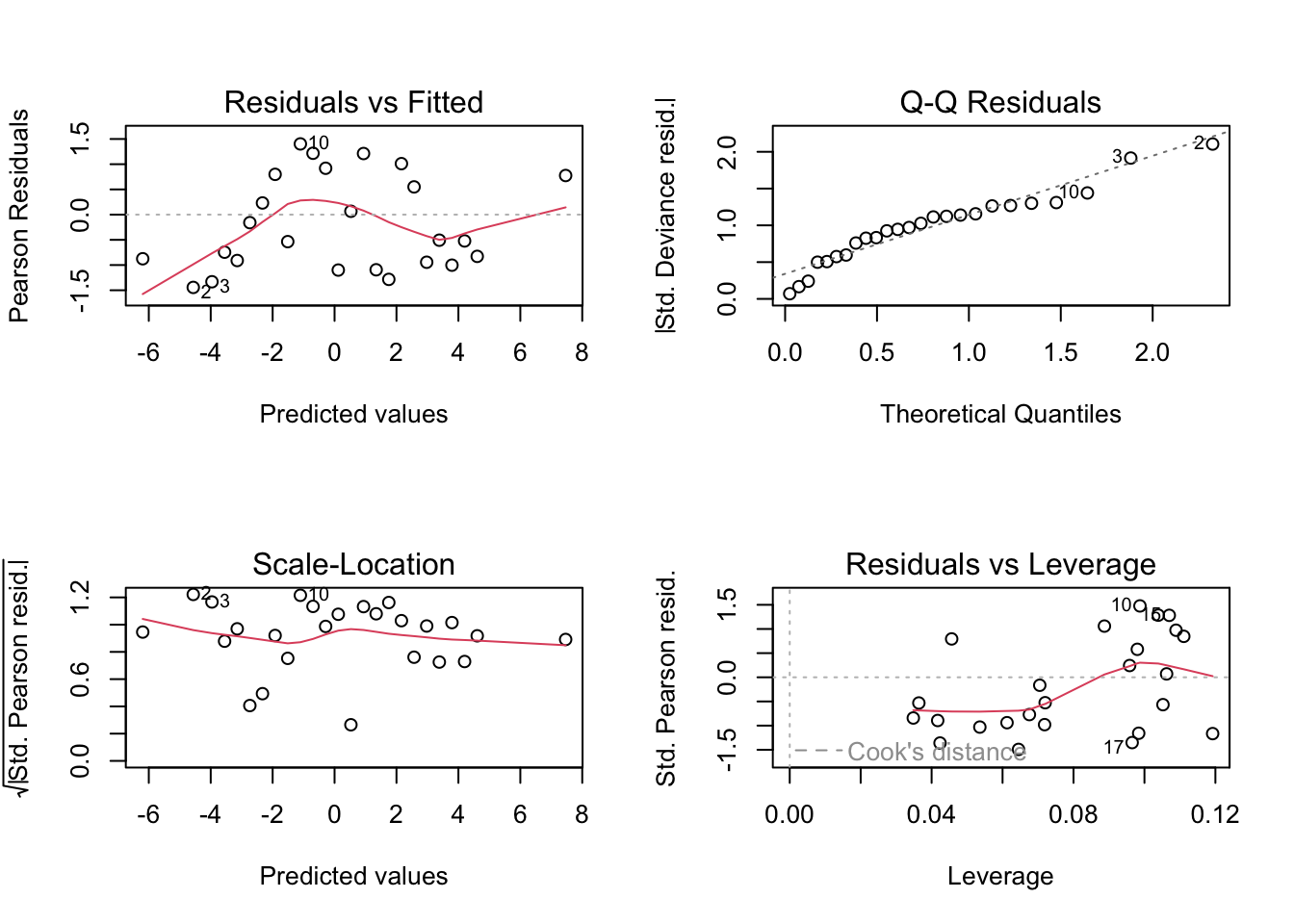
Thus, we obtain very similar results using PCR or PLS in this case.
The fundamental difference between PCR and PLS is in the way they incorporate the \(y\) values into the model. PLS does this directly by involving covariances between \(y\) and all the \(x_i\) variables. PCR derives its “factors” independently of \(y\), then makes the link by straightforward multiple regression. One consequence of this difference seems to be that to achieve the same level of predictive ability PCR models generally require one, sometimes two, more components than the corresponding PLS model. For both PCR and PLS, the number of terms for the model should be determined by cross-validation. The function plsr in the pls package does this for us. We have not used cross-validation for the PCR example above simply because it would involve some messy R programming to it by hand. In fact the pls package includes a pcr function which does do cross-validation for PCR as for PLS, but that function does not incorporate the flexibility of selecting your PCs though. It forces you to use the first \(k\) PCs, which as we have seen from our example, is not always the most sensible choice.
In both the PCR and PLS modelling of the NIR data we did not standardise the \(X\) values, the NIR wavelengths, because they are all measured on the same scale. It is more common, when the \(X\) variables are measured on disparate scales, that we standardise by centring (subtract the mean) and scale (divide by the standard deviation). The argument scale does this in the pls function. However one must be careful with this operation in high-dimensional data as it would give the same relevance to genuine and noisy signals.
Finally note that, as with PCR, we can convert the PLS model back to the original wavelengths. The R code is similar and the resulting PLS model (not shown) is almost identical to the PCR model.