Chapter 6 Multidimensional scaling
6.1 Introduction
We have noted that principal components analysis produces a low-dimensional representation of the data, typically in two dimensions to facilitate interpretation. The relative position of the observations or samples in this representation approximates the similarity between them according to the measured variables. For ordinary numerical data this spatial configuration approximates the Euclidean distance between the original samples. This chapter introduces a class of methods that aim to produce an analogous low-dimensional representation from a distance or similarity matrix.
This provides a more general and fit-for-purpose approach to undergo analysis of similarities between samples as it allows to:
- Use measures other than the Euclidean distance, actually whichever is regarded suitable for the nature of the data.
- Work directly with a distance or similarity matrix, i.e. one obtained as raw data and not one derived from an ordinary data set \(\mathbf X\) consisting of observations for a number variables. For example, data obtained from judges assessing how alike a number of cars, wines, tissue lesions, etc. are.
Note that if working with numerical data and using the Euclidean distance then the configuration of points will coincide with that obtained using PCA. This stresses the idea that many multivariate methods are not exclusive, they can be used in conjunction, offering different insight into the same data from different viewpoints. These methods are known as ordination methods in ecological and environmental sciences, in reference to ordering samples along axes of scientific meaning, but they are also widely used in other disciplines,
Starting with a distance or similarity matrix, the aim is to find a small number of new axes, or dimensions, which explain most of the distance or similarity matrix information. We then plot the samples along these new axes and observe their new orderings or ordinations.
An intuitively appealing example of this process uses, as its starting point, the inter-town distances table often included in old-fashioned road atlases. Treating this as a distance matrix, we generate two axes from it such that the towns (samples), when plotted on these new axes, are approximately the same distances apart as those given in the original matrix. In this case, because the original distances are almost Euclidean, the first two new axes will account for almost all the distance information and our “map” of the towns will be quite accurate. However, there is indeterminacy in terms of orientation. Our new map might be upside down or run East – West instead of West – East. The original matrix of distances contains no information on this point so neither do the new ordinations.
The main output is a low-dimensional graphical representation where the spatial distribution of the points optimally approximates the relative similarity between the samples. There are several methods to produce this and they largely differ in how agreement between fitted distances and observed proximities is assessed. We focuses here on classical or metric multidimensional scaling (MDS), also known as principal coordinates analysis.
6.2 Basic elements
Given a distance matrix \({d_{ij}}\) for \(n\) samples the aim is to find the best \((n-1)\)-column matrix of coordinates to describe the \({d_{ij}}\), with “best” here meaning that the (Euclidean) distances \(\delta_{ij}\) derived from the matrix of coordinates minimise \[\psi = \sum (d^{2}_{ij}-\delta^{2}_{ij})\] Each column of the matrix form an axis or dimension providing the scores for each sample. It is desirable that the first 2-3 axes account for most of the distance information, so that we can obtain a succinct 2D or 3D graphical summary of the original (sometimes large) similarity/distance matrix and facilitate the interpretation of the results. As for the previous techniques, the axes are uncorrelated and can be ordered so that they successively account for decreasing fractions of the original information. Note that in MDS we only have scores and not loadings, we work directly with the distance matrix and have no information about the original variables.
For interpretation it is useful to plot the scores labelled by factors of interest or against other relevant variables.
6.3 MDS of the moths data set
The base R function implementing MDS is cmdscale. We use the normalised Manhattan distance matrix of the logged species counts created previously and compute \(k = 2\) MDS dimensions or axes.
# From previous chapter:
moths <- read.table("moths.txt", h = T, stringsAsFactors = TRUE)
mothslog <- moths # Create a separate data set
mothslog[, 5:16] <- log10(mothslog[, 5:16] + 1) # Log-transform the counts
mothslog$Type <- as.factor(mothslog$Type) # To be used later
# Normalised Manhattan distance matrix using vegan package
library(vegan)
manhdist <- vegdist(mothslog[, 5:16], method = "gower")The results are held as follows:
- $points: coordinates for the MDS axes.
- $eig: contributions of each dimension to the total distance information provided by eigenvalues (similar to % variance due to each principal component in PCA).
- $GOF: goodness of fit, % distance accounted for by all k dimensions considered.
We can visualise the individual dimension contributions to represent the distances (eigenvalues, eig, are computed for all possible MDS axes, 14 in this case).
What is the goodness of fit (GOF) if we reduce to k dimensions? The value of the sum of the first \(k\) eigenvalues with respect to the total sum of eigenvalues could be used as for PCA, but this involves technical problems if no Euclidean distance is used and negative eigenvalues are produced. Two alternative measures are provided by the cmdscale function:
\[GOF=\frac{\sum_{i=1}^k|\lambda_i|}{\sum_{i=1}^n|\lambda_i|} \;\;\mbox{and}\;\; GOF=\frac{\sum_{i=1}^k\max(0,\lambda_i)}{\sum_{i=1}^n\max(0,\lambda_i)}\] Note that these two measures are equal when Euclidean distance is used.
[1] 0.6457675 0.7020417The first of these measures suggests that 65% of the distance information is accounted for by the first 2 MDS axes. In fact, the plot above suggests that perhaps using up to four.
The following scatterplot shows the scores using the first two MDS axes. The points are labelled by site type.
colours <- c("black", "red", "green", "blue")[mothslog$Type]
plot(mds$points[, 1:2], col = colours, xlab = "MDS1", ylab = "MDS2")
legend("topright", legend = levels(mothslog$Type), col = 1:3, pch = 1)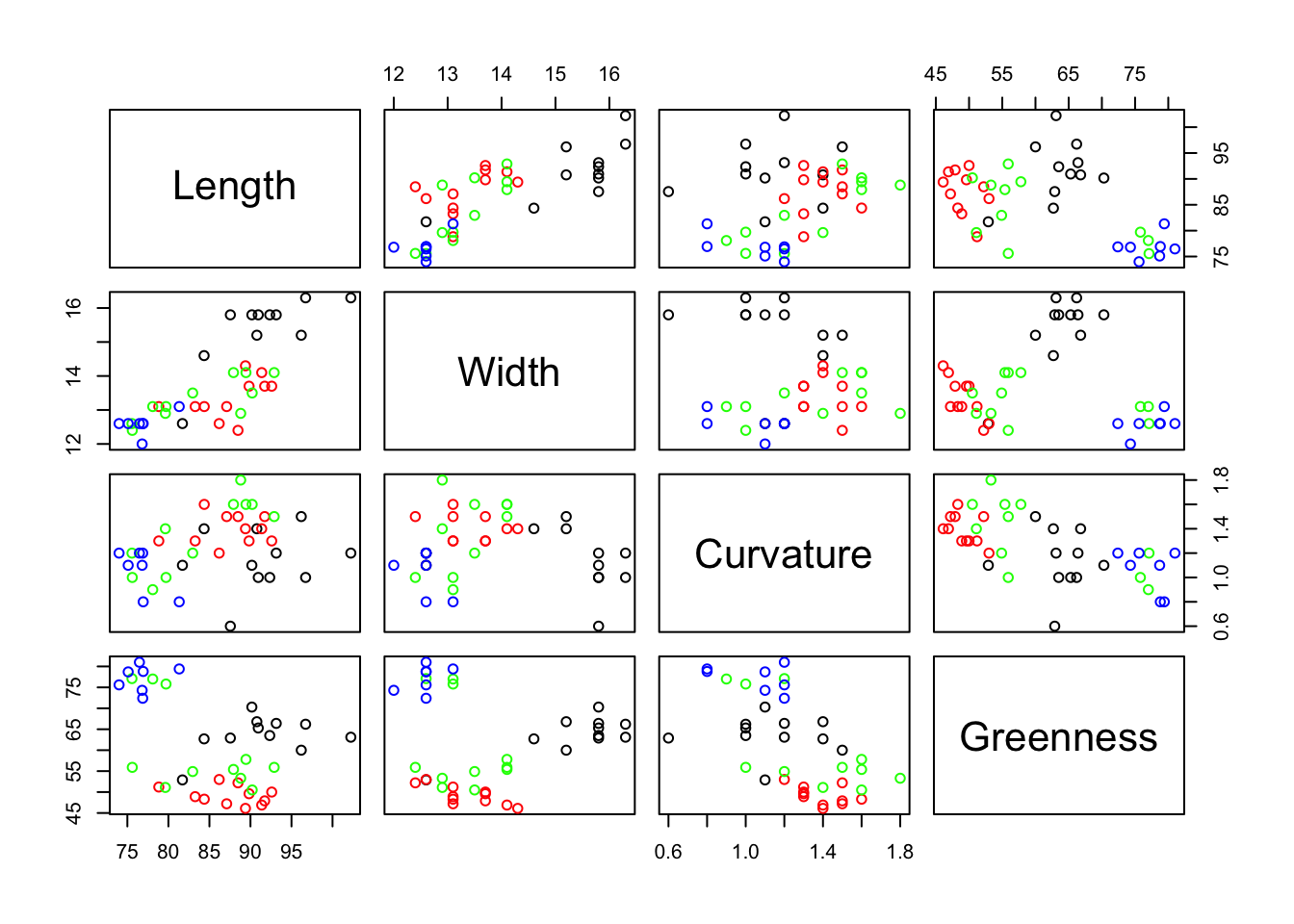
The plot above suggests that the 3 Farmland sites (in black, F) have similar moths profiles as they are close to each other, but there do not seem to be any other patterns.
However, we do have other variables in the data set, the Northings and Eastings and these can be easily incorporated into the graph.
# Plot scores with no symbol
plot(mds$points[, 1:2], type = "n", xlab = "MDS1", ylab = "MDS2")
# Use the Northings coordinates to label them instead
text(mds$points[, 1], mds$points[, 2], labels = mothslog$N)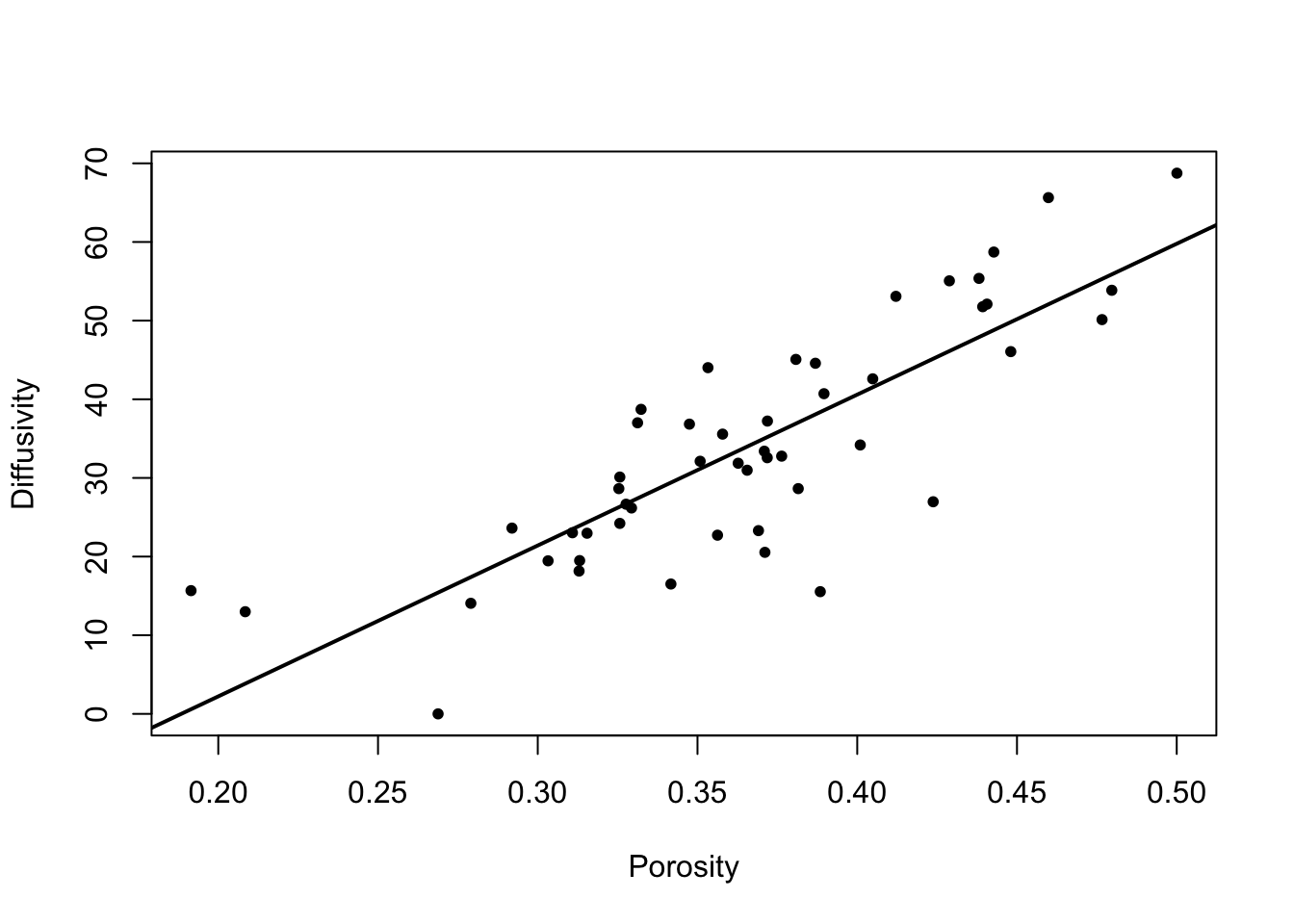
This graph indicates that the first MDS dimension is mainly a North-South gradient.
6.4 Exercises
RAPD data. 26 nematode (tiny worm which lives in soil) samples representing 5 species were assessed on 122 RAPD bands (one of the early genetic marker technologies). Each band can take the value 1 or 0. Using the Jaccard coefficient derive a multidimensional scaling plot for the 25 samples. Do they separate into their known species? How many MDS dimensions are needed?
LIFEEXP data. Does a MDS analysis of these data provide a succinct summary of the similarities and differences among the countries with respect to life expectancy?